
- #Deleting base pairs in bioedit manual#
- #Deleting base pairs in bioedit full#
- #Deleting base pairs in bioedit software#
- #Deleting base pairs in bioedit trial#
- #Deleting base pairs in bioedit free#
One quirk of BioEdit is that if you double click a data file it will open in a new copy of BioEdit, not in an existing one. I would recommend saving everything in fasta format since that is the format I use in order to convert the data to another format or to another person (who probably doesn’t have a copy of BioEdit). As far as I can tell there is no difference between saving your file as a BioEdit formatted file versus as a fasta file. I use this feature on nearly every dataset I create. It is the only program I know of that allows you to edit, search and replace, and paste over the sequence title names independent of your sequences. Aside from that limitation (which isn’t as bad as it might sound once you learn a few tricks), I really like its features. The most annoying aspect is that you have to manually align up each sequence and manually create a consensus sequence (which commercial programs like Sequencher and Geneious are very good at).
#Deleting base pairs in bioedit free#
I use BioEdit to align sequences as it is free and has some handy features.
#Deleting base pairs in bioedit trial#
Each of the commercial versions have a free 60 day trial should you wish to try them. I’ve always used the free Chromas version, Chromas Lite, but there are two other versions with more features that are fairly cheap. One trick I find useful later is to always edit your sequences from the same starting base (unless the starts are all messy), as it makes sequence alignment much easier later. Alternatively, you can go edit, copy sequence, FASTA format and paste that into BioEdit. When I am done I save the chromatogram and export the data to a line file (which is saved with a. I usually make all of my edits as lower case bases as it makes it easier to identify where I have made edits. The chromatograms come off the machine with all bases in upper case. This opens the file in Chromas (see below under installation notes if some other program opens it instead of Chromas). MEGA also has an alignment editor, but I’ve not really used it very much.ĭouble click on the chromatogram file (usually has the extension ab1). BioEdit can also edit chromatograms, but I find Chromas to be nicer.

If I loose my sequence alignment, at least all my chromatograms with the correct edits are still there to rebuild it from. Our experiments indicate that estimates of population mutation rates can be affected two- to three-fold by uncorrected errors.Guide to editing sequences with Chromas and BioEdit Guide to editing sequences with Chromas and BioEditīy Peter Unmack Editing chromatograms with ChromasĬhromas has the advantage the you can save all of your chromatograms which can subsequently be used in any other programs unlike Sequencher which saves everything in a project file which cannot be opened by anything else.
Finally, we show that correcting sequencing errors is important, because population genetic and phylogenetic inferences can be misled by MSAs with uncorrected mis-calls. After this pre-filtering, the user only needs to inspect a small number of peaks in every chromatogram to correct sequencing errors. The threshold value represents the sensitivity of the sequencing error detection mechanism. The guided MSA assembly procedure in ChromatoGate detects chromatogram peaks of all characters in an alignment that lead to polymorphic sites, given a user-defined threshold. Initially, the program scans a given multiple sequence alignment (MSA) for potential sequencing errors, assuming that each polymorphic site in the alignment may be attributed to a sequencing error with a certain probability.
#Deleting base pairs in bioedit full#
To provide users full control over the error correction process, a fully automated error correction algorithm has not been implemented.
#Deleting base pairs in bioedit software#
Here, we introduce ChromatoGate (CG), an open-source software that accelerates and partially automates the inspection of chromatograms and the detection of sequencing errors for bidirectional sequencing runs.
#Deleting base pairs in bioedit manual#
As sequence numbers and lengths increase, visual inspection and manual correction of chromatograms and corresponding sequences on a per-peak and per-nucleotide basis becomes an error-prone, time-consuming, and tedious process. Since chromatogram translation programs frequently introduce errors, a manual inspection of the generated sequence data is required.
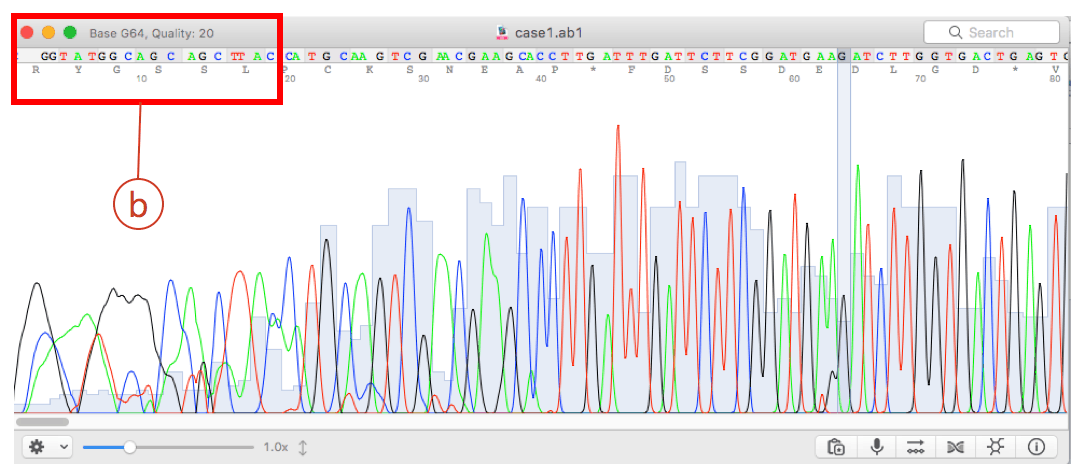
They also generate data that translates the chromatograms into molecular sequences of A, C, G, T, or N (undetermined) characters. Automated DNA sequencers generate chromatograms that contain raw sequencing data.

